QCoDeS example with NanoVNA H4¶
The NanoVNA is a USB-controlled vector network analyser. This driver was tested for version H4. ## Set up the instrument
Make sure you have QCoDeS set up (see the QCoDeS website or my notebook 14 minutes to QCoDeS).
In your
qcodesenvironment, install pynanovna.Plug the instrument into the USB interface of your computer and turn it on. A good testing configuration is to connect ports 1 and 2 via an SLP-100 filter.
Run the following code, and check that you get a connect message.
[1]:
import qcodes as qc
from qcodes_contrib_drivers.drivers.NanoVNA.H4 import NanoVNA
qc.Instrument.close_all() # Closes all open instruments, in case there is a duplicate for any reason.
vna = NanoVNA("vna") # Should report something like: Connected to: NanoVNA NanoVNA (serial:None, firmware:1.2.27) in 1.94s
2026-02-11 19:30:28,564 - root - INFO - Initializing the VNA.
2026-02-11 19:30:28,864 - pynanovna.hardware.Hardware - INFO - Finding firmware variant...
2026-02-11 19:30:29,876 - root - INFO - VNA successfully initialized.
2026-02-11 19:30:30,296 - pynanovna.hardware.Hardware - INFO - Finding firmware variant...
2026-02-11 19:30:30,421 - qcodes.instrument.instrument_base - INFO - [vna(NanoVNA)] Connected to instrument: {'vendor': 'NanoVNA', 'model': 'NanoVNA', 'serial': None, 'firmware': '1.2.27'}
Connected to: NanoVNA NanoVNA (serial:None, firmware:1.2.27) in 1.86s
Initialise QCoDeS control¶
[2]:
from qcodes.station import Station
from qcodes.dataset import (
do0d,
do1d,
do2d,
initialise_database,
initialise_or_create_database_at,
load_or_create_experiment,
Measurement,
plot_by_id
)
# create an empty database based on the config file
qc.initialise_or_create_database_at('./test_vna.db')
exp = load_or_create_experiment(experiment_name='testing_NanoVNA',
sample_name="low_pass_filter")
station = qc.Station(vna)
2026-02-11 19:30:30,861 - pynanovna.hardware.Hardware - INFO - Finding firmware variant...
2026-02-11 19:30:31,483 - root - CRITICAL - No calibration has been applied, it is strongly recommended to calibrate you NanoVNA.
2026-02-11 19:30:31,485 - root - CRITICAL - 1 port calibration not valid, it is recommended to re-calibrate.
2026-02-11 19:30:31,487 - root - CRITICAL - 2 port calibration not valid, it is recommended to re-calibrate.
In my setup, I get the error 634 - root - CRITICAL - No calibration has been applied, it is strongly recommended to calibrate you NanoVNA. This happens even when I have manually calibrated the instrument; I don’t know how to avoid this. ## Connect to the instrument and make some measurements
[3]:
vna.start_freq(1e6)
vna.stop_freq(900e6)
vna.npts(301)
meas = Measurement()
meas.register_parameter(vna.frequency)
meas.register_parameter(vna.s11_real)
meas.register_parameter(vna.s11_imag)
meas.register_parameter(vna.s11_mag_db)
meas.register_parameter(vna.s11_mag_lin)
meas.register_parameter(vna.s11_phase)
meas.register_parameter(vna.s21_real)
meas.register_parameter(vna.s21_imag)
meas.register_parameter(vna.s21_mag_db)
meas.register_parameter(vna.s21_mag_lin)
meas.register_parameter(vna.s21_phase)
with meas.run() as datasaver:
datasaver.add_result(
(vna.frequency, vna.frequency()),
(vna.s11_real, vna.s11_real()),
(vna.s11_imag, vna.s11_imag()),
(vna.s11_mag_db, vna.s11_mag_db()),
(vna.s11_mag_lin, vna.s11_mag_lin()),
(vna.s11_phase, vna.s11_phase()),
(vna.s21_real, vna.s21_real()),
(vna.s21_imag, vna.s21_imag()),
(vna.s21_mag_db, vna.s21_mag_db()),
(vna.s21_mag_lin, vna.s21_mag_lin()),
(vna.s21_phase, vna.s21_phase()),
)
plot_by_id(datasaver.run_id)
2026-02-11 19:30:31,496 - qcodes.dataset.measurements - INFO - Registered vna_frequency in the Measurement.
2026-02-11 19:30:31,497 - qcodes.dataset.measurements - INFO - Registered vna_s11_real in the Measurement.
2026-02-11 19:30:31,498 - qcodes.dataset.measurements - INFO - Registered vna_s11_imag in the Measurement.
2026-02-11 19:30:31,499 - qcodes.dataset.measurements - INFO - Registered vna_s11_mag_db in the Measurement.
2026-02-11 19:30:31,499 - qcodes.dataset.measurements - INFO - Registered vna_s11_mag_lin in the Measurement.
2026-02-11 19:30:31,500 - qcodes.dataset.measurements - INFO - Registered vna_s11_phase in the Measurement.
2026-02-11 19:30:31,501 - qcodes.dataset.measurements - INFO - Registered vna_s21_real in the Measurement.
2026-02-11 19:30:31,501 - qcodes.dataset.measurements - INFO - Registered vna_s21_imag in the Measurement.
2026-02-11 19:30:31,502 - qcodes.dataset.measurements - INFO - Registered vna_s21_mag_db in the Measurement.
2026-02-11 19:30:31,503 - qcodes.dataset.measurements - INFO - Registered vna_s21_mag_lin in the Measurement.
2026-02-11 19:30:31,504 - qcodes.dataset.measurements - INFO - Registered vna_s21_phase in the Measurement.
2026-02-11 19:30:31,516 - qcodes.dataset.sqlite.queries - INFO - Created run with guid: a554923d-0000-0000-0000-019c4e2f09d2 in database C:\Users\lairde\OneDrive - Lancaster University\Desktop\Sandpit\test_vna.db
2026-02-11 19:30:31,552 - qcodes.dataset.sqlite.queries - INFO - Set the run_timestamp of run_id 15 to 1770838231.5522954
2026-02-11 19:30:31,556 - qcodes.dataset.measurements - INFO - Starting measurement with guid: a554923d-0000-0000-0000-019c4e2f09d2, sample_name: "low_pass_filter", exp_name: "testing_NanoVNA", ds_name: "results".
2026-02-11 19:30:31,557 - qcodes.dataset.measurements - INFO - Using background writing: False
Starting experimental run with id: 15.
2026-02-11 19:30:32,444 - root - CRITICAL - No calibration has been applied, it is strongly recommended to calibrate you NanoVNA.
2026-02-11 19:30:32,445 - root - CRITICAL - 1 port calibration not valid, it is recommended to re-calibrate.
2026-02-11 19:30:32,445 - root - CRITICAL - 2 port calibration not valid, it is recommended to re-calibrate.
2026-02-11 19:30:32,464 - qcodes.dataset.measurements - INFO - Finished measurement with guid: a554923d-0000-0000-0000-019c4e2f09d2.
[3]:
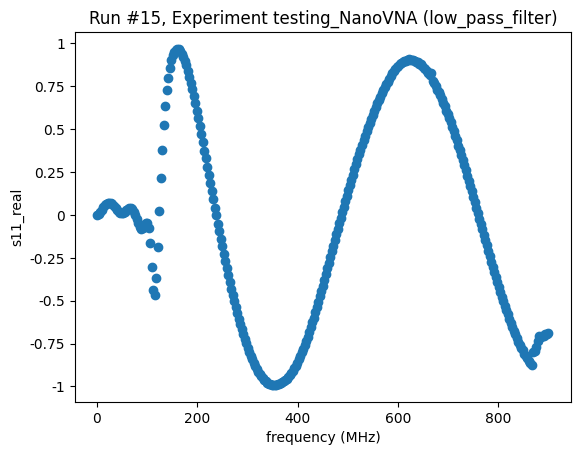
([<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s11_real'>,
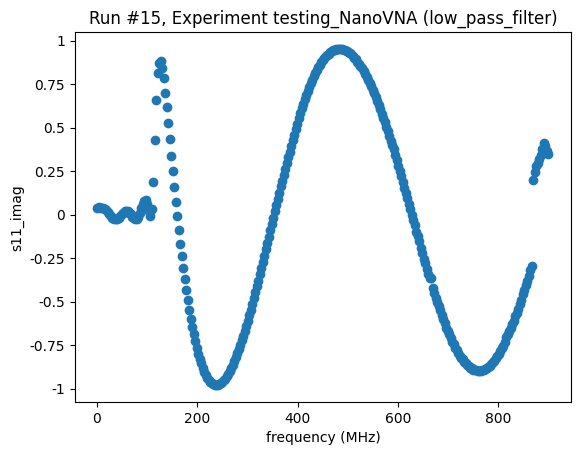
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s11_imag'>,
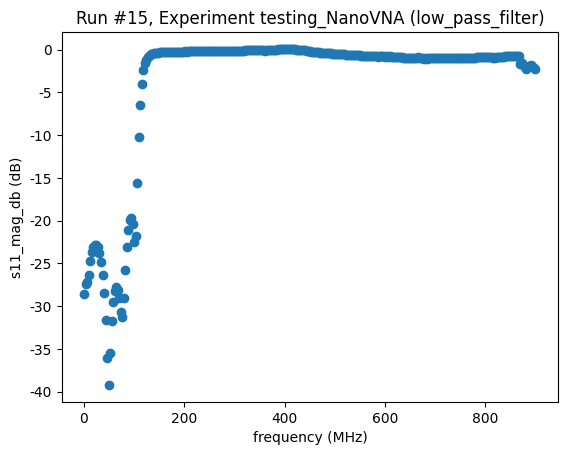
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s11_mag_db (dB)'>,
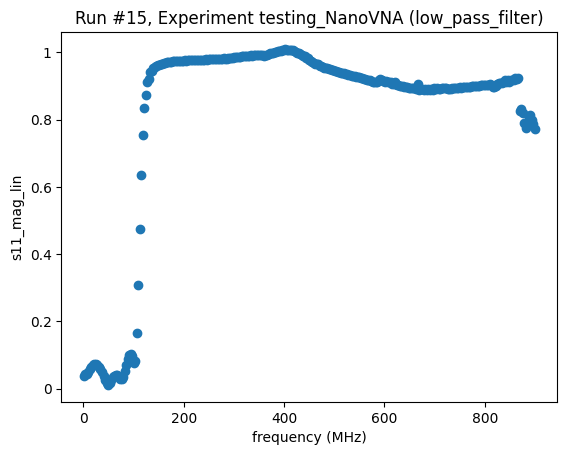
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s11_mag_lin'>,
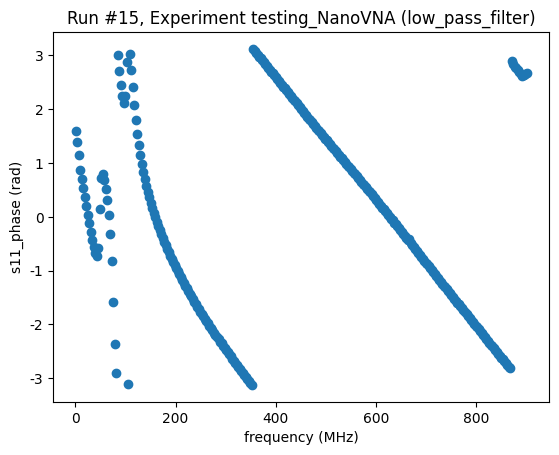
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s11_phase (rad)'>,
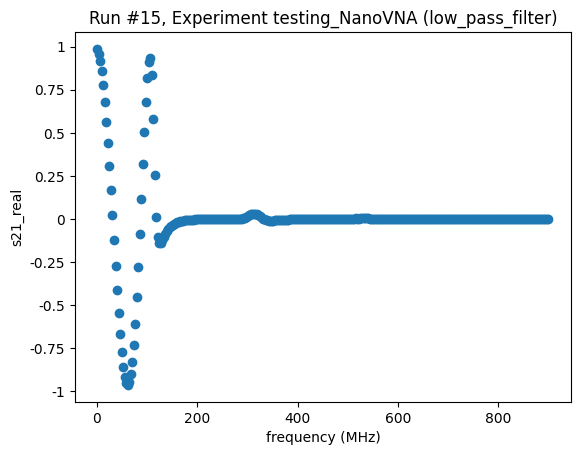
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s21_real'>,
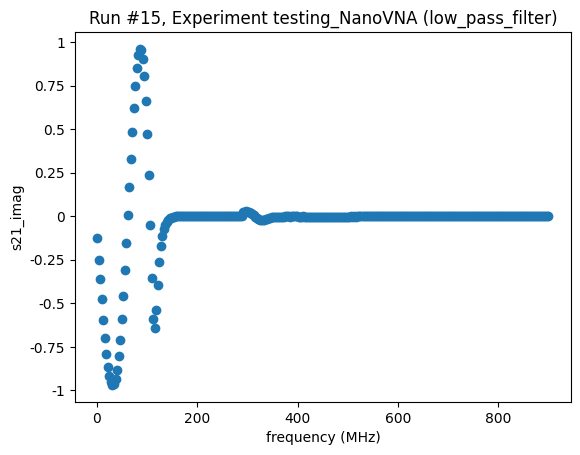
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s21_imag'>,
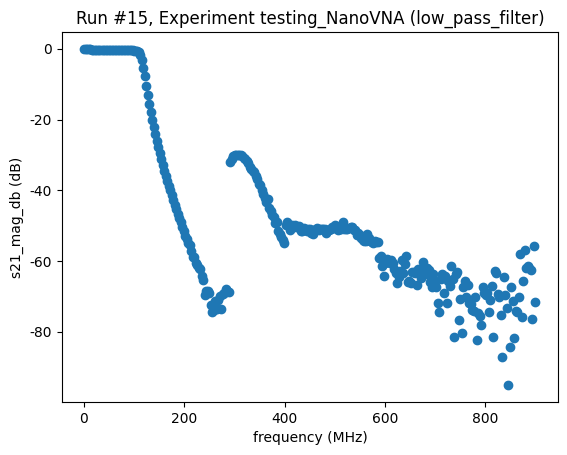
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s21_mag_db (dB)'>,
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s21_mag_lin'>,
<Axes: title={'center': 'Run #15, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s21_phase (rad)'>],
[None, None, None, None, None, None, None, None, None, None])
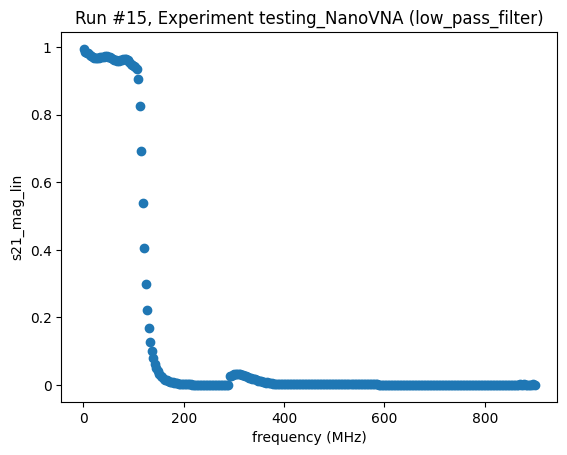
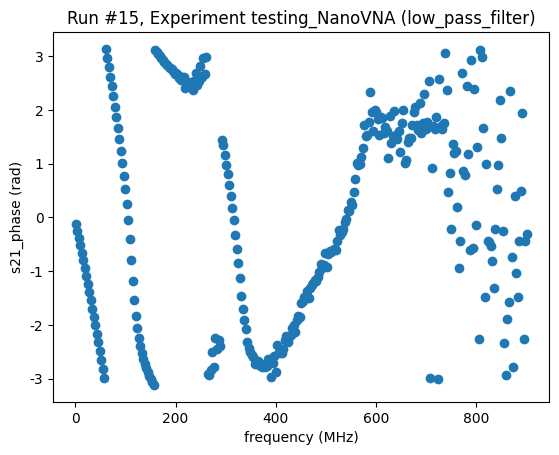
Do a QCoDeS sweep, measuring the magnitudes of S11 and S12¶
[4]:
do0d(vna.s11_mag_db, vna.s21_mag_db, do_plot=True)
2026-02-11 19:30:33,185 - qcodes.dataset.measurements - INFO - Registered vna_s11_mag_db in the Measurement.
2026-02-11 19:30:33,186 - qcodes.dataset.measurements - INFO - Registered vna_frequency in the Measurement.
2026-02-11 19:30:33,187 - qcodes.dataset.measurements - INFO - Registered vna_s21_mag_db in the Measurement.
2026-02-11 19:30:33,187 - qcodes.dataset.dond.do_nd - INFO - Starting a doNd with scan with
setpoints: (),
measuring: (<qcodes.parameters.parameter_with_setpoints.ParameterWithSetpoints: s11_mag_db at 2735099484816>, <qcodes.parameters.parameter_with_setpoints.ParameterWithSetpoints: s21_mag_db at 2735099486736>)
2026-02-11 19:30:33,196 - qcodes.dataset.sqlite.queries - INFO - Created run with guid: d3f24f1f-0000-0000-0000-019c4e2f1065 in database C:\Users\lairde\OneDrive - Lancaster University\Desktop\Sandpit\test_vna.db
2026-02-11 19:30:33,212 - qcodes.dataset.sqlite.queries - INFO - Set the run_timestamp of run_id 16 to 1770838233.2120826
2026-02-11 19:30:33,215 - qcodes.dataset.measurements - INFO - Starting measurement with guid: d3f24f1f-0000-0000-0000-019c4e2f1065, sample_name: "low_pass_filter", exp_name: "testing_NanoVNA", ds_name: "results". Using 'qcodes.dataset.do0d'
2026-02-11 19:30:33,215 - qcodes.dataset.measurements - INFO - Using background writing: False
2026-02-11 19:30:33,230 - qcodes.dataset.measurements - INFO - Finished measurement with guid: d3f24f1f-0000-0000-0000-019c4e2f1065. Using 'qcodes.dataset.do0d'
Starting experimental run with id: 16. Using 'qcodes.dataset.do0d'
[4]:
(results #16@C:\Users\lairde\OneDrive - Lancaster University\Desktop\Sandpit\test_vna.db
---------------------------------------------------------------------------------------
vna_frequency - array
vna_s11_mag_db - array
vna_s21_mag_db - array,
(<Axes: title={'center': 'Run #16, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s11_mag_db (dB)'>,
<Axes: title={'center': 'Run #16, Experiment testing_NanoVNA (low_pass_filter)'}, xlabel='frequency (MHz)', ylabel='s21_mag_db (dB)'>),
(None, None))
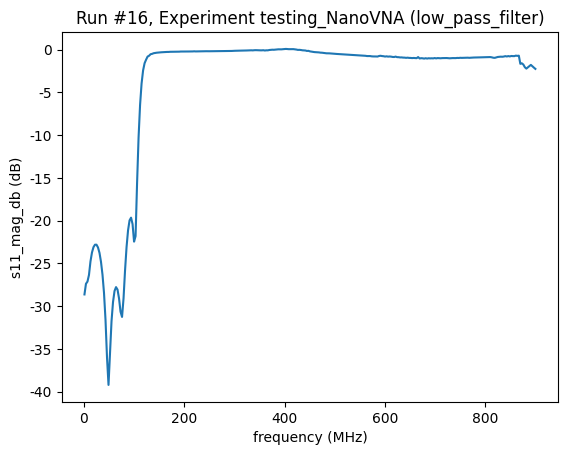
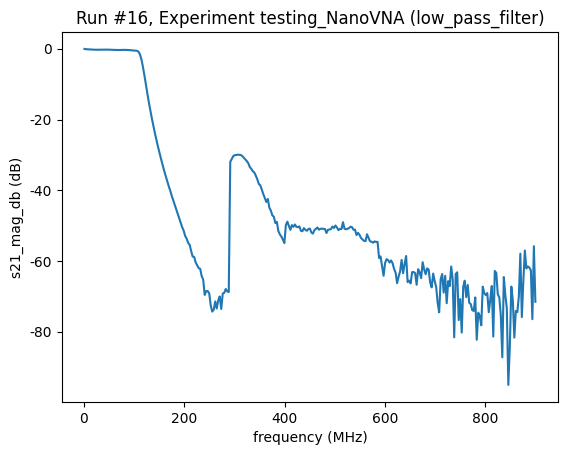
Close the connection¶
[5]:
vna.close()
2026-02-11 19:28:59,932 - pynanovna.hardware.VNABase - INFO - disconnect COM4 (NanoVNA)